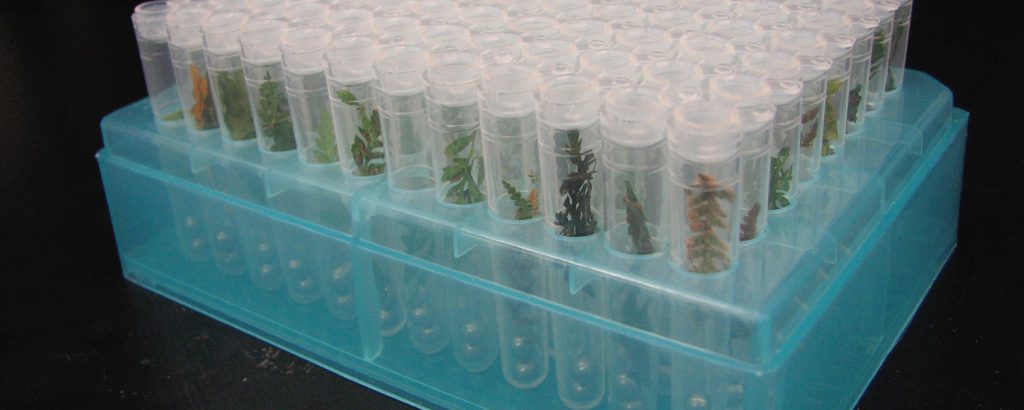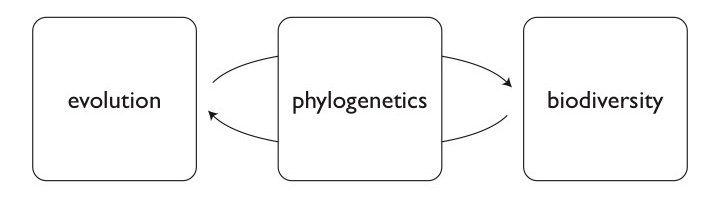
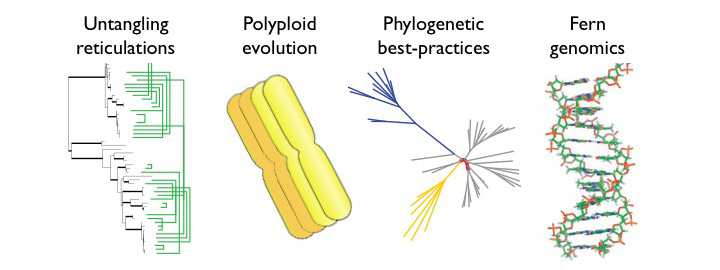
We use the tools of phylogenetics to explore the process of evolution and the patterns it creates. Research in the lab focuses on (but is not restricted to) ferns and lycophytes, with particular emphasis on 1) developing and applying phylogenetic tools to understand reticulate evolution (evolution that involves hybridization as well as branching of lineages); 2) inferring the evolutionary fate of polyploids (those taxa that have more than two copies of all of their chromosomes); 3) understanding the strengths and weakness of our phylogenetic methods; and 4) developing new genomic tools to better study poorly-understood groups (like ferns!).
Polyploid Evolution
Are polyploids dead-ends, on evolutionary timescales? Given that polyploid lineages are common and often ecologically highly successful, this may seem like a silly question. However, there is not necessarily a contradiction here–instead of polyploids themselves diversifying (the scenario on the right, below) polyploids could be common because they form frequently (even if they subsequently tend to go extinct quickly; the scenario on the left) and they could enjoy short-term successes yet still be prone to longer-term evolutionary failure.
Reticulate evolution
One recent project has been to develop a web lab workflow and associated bioinformatic pipeline–“PURC“–to generate phylogenetic data for polyploids, which we hope will facilitate research into a wide variety of understudied polyploid complexes.
![]()
Cystopteridaceae and the Cystopteris fragilis complex
One of the main taxonomic groups that we focus on is Cystopteridaceae, the ecologically diverse and globally distrbuted fern family that includes Cystopteris (the fragile ferns, bladder ferns, and bulblet fern), Acystopteris, Gymnocarpium (the oak ferns), and xCystocarpium (the Cystopteris–Gymnocarpium hybrid).
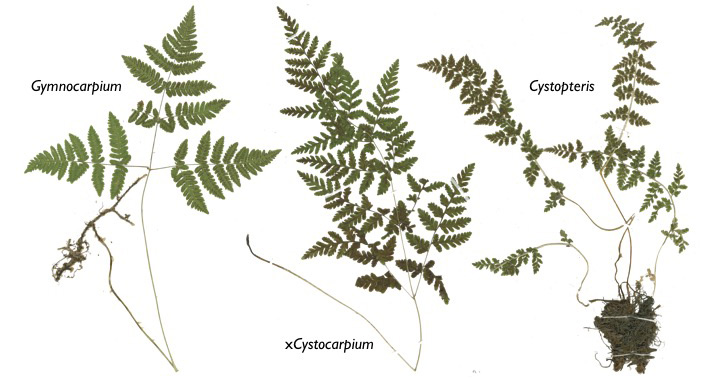
Cystopteridaceae contains a gloriously distressing array of polyploid complexes with taxa ranging from diploid (two chromosome sets) to octaploid (eight sets), particularly in the infamous “Cystopteris fragilis complex.”
Research Approaches